Functional Immunogenomics
TEL +81-6-6879-4926
Overview
Each cell in the human body contains the same genetic material, yet the nuanced use of this shared genetic code underlies the vast diversity of cell types and functions we observe. Recent advances in genetic engineering tools and single cell genomics technologies have enabled an unprecedented opportunity to study these regulatory interactions in depth and at scale (Figure 1).
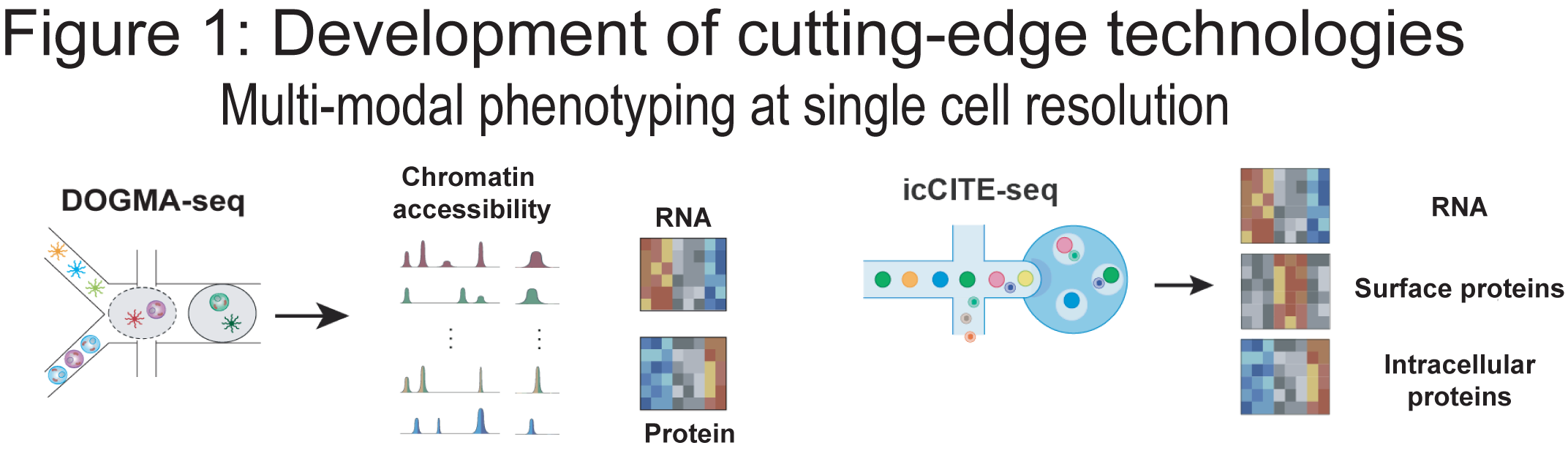
Our research aims to understand how gene regulatory programs control immune cell fate and function. By integrating perturbation-based screens with multi-modal single-cell profiling, we systematically dissect the genetic and epigenetic mechanisms that govern cell state, differentiation and function (Figure 2). In parallel, we apply these cutting-edge approaches to profile patient-derived samples, enabling us to characterize immune dysregulation in disease contexts and link regulatory mechanisms to relevant cell states (Figure 2).
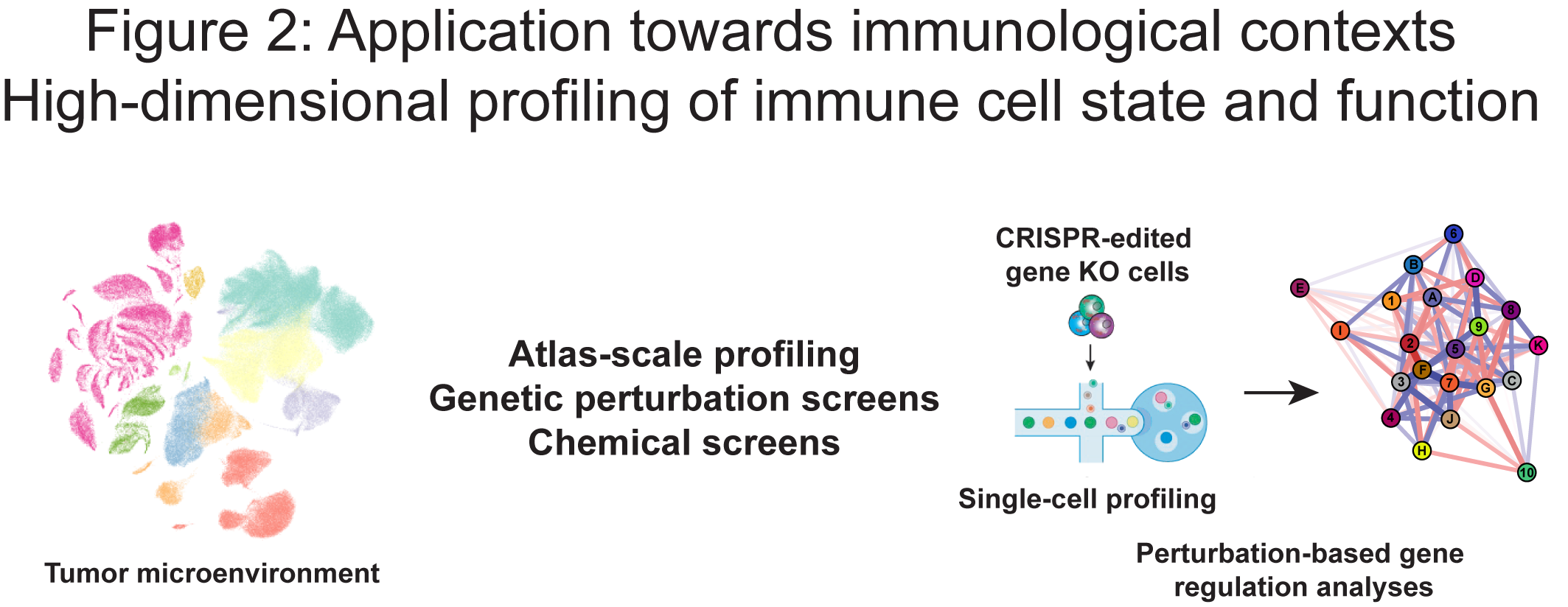
Through these complementary strategies, we seek to identify key regulatory elements and pathways that can be harnessed to modulate immune responses. Ultimately, our goal is to translate these insights into next-generation adoptive cell therapies for the treatment of diseases.
Principal Investigator
Kelvin Chen Associate Professor

Research field
Immunology, Immunogenomics, Gene regulation
Education history
| 2014 | Bachelor of Science, Biological Sciences, The University of Osaka |
| 2016 | Master of Medical Sciences, The University of Osaka |
| 2021 | Doctor of Philosophy, Medicine, The University of Osaka |
Research and career history
| 2014 | Research assistant, IFReC, The University of Osaka |
| 2020 | Researcher, Institute for Frontier Life and Medical Sciences, Kyoto University |
| 2021 | Researcher, IFReC, The University of Osaka |
| 2021 | Specially-appointed Assistant Professor, IFReC, The University of Osaka |
| 2026 | Associate Professor, IFReC, The University of Osaka |
Prize
| 2025 | Osaka University Prize |
Members
- Kelvin Chen Associate Professor
chenkifrec.osaka-u.ac.jp
Achievements
Publications
- Mikami N, Kawakami R, Chen KY, Sugimoto A, Ohkura N, Sakaguchi S. Epigenetic conversion of conventional T cells into regulatory T cells by CD28 signal deprivation. Proc Natl Acad Sci U S A. 2020 Jun 2;117(22):12258–68.
- Kawakami R, Kitagawa Y, Chen KY, Arai M, Ohara D, Nakamura Y, Yasuda K, Osaki M, Mikami N, Lareau CA, Watanabe H, Kondoh G, Hirota K, Ohkura N, Sakaguchi S. Distinct Foxp3 enhancer elements coordinate development, maintenance, and function of regulatory T cells. Immunity. 2021 May 11;54(5):947–61.e8.
- Mimitou EP, Lareau CA, Chen KY, Zorzetto-Fernandes AL, Hao Y, Takeshima Y, Luo W, Huan TS, Yeung BZ, Paplexi E, Thakore PI, Kibayashi T, Wing JB, Hata M, Satija R, Nazor KL, Sakaguchi S, Ludwig LS, Sankaran VG, Regev A, Smibert P. Scalable, multimodal profiling of chromatin accessibility, gene expression and protein levels in single cells. Nat Biotechnol. 2021 Oct;39(10):1246–58.
- Chen KY, Kibayashi T, Giguelay A, Hata M, Nakajima S, Mikami N, Takeshima Y, Ichiyama K, Omiya R, Ludwig LS, Hattori K, Sakaguchi S. Genome-wide CRISPR screen in human T cells reveals regulators of FOXP3. Nature. 2025 Mar 26;642(8066):191–200.
- Arase M, Murakami M, Kihara T, Kuwahara R, Toyota H, Sumitani N, Kinoshita N, Chen KY, Yokoi T, Motooka D, Okuzaki D, Zhao Y, Miyazaki H, Ogino T, Hirota S, Ikeuchi H, Takeda K. Multi-omics uncovers transcriptional programs of gut-resident memory CD4+ T cells in Crohn’s disease. J Exp Med [Internet]. 2025 Nov 3;222(11).
